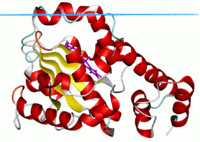
CRAL-TRIO domain
| CRAL/TRIO domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
|
Alpha-tocopherol transfer protein, closed state with ligand.[1] | |||||||||
| Identifiers | |||||||||
| Symbol | CRAL_TRIO | ||||||||
| Pfam | PF00650 | ||||||||
| InterPro | IPR001251 | ||||||||
| SMART | Sec14 | ||||||||
| SCOP | 1aua | ||||||||
| SUPERFAMILY | 1aua | ||||||||
| OPM superfamily | 129 | ||||||||
| OPM protein | 1r5l | ||||||||
| CDD | cd00170 | ||||||||
| |||||||||
CRAL-TRIO domain is a protein structural domain that binds small lipophilic molecules.[2] This domain is named after cellular retinaldehyde-binding protein (CRALBP) and TRIO guanine exchange factor.
CRALB protein carries 11-cis-retinol or 11-cis-retinaldehyde. It modulates interaction of retinoids with visual cycle enzymes. TRIO is involved in coordinating actin remodeling, which is necessary for cell migration and growth.
Other members of the family are alpha-tocopherol transfer protein and phosphatidylinositol-transfer protein (Sec14). They transport their substrates (alpha-tocopherol and phosphatidylinositol or phosphatidylcholine, respectively) between different intracellular membranes. Family also include a guanine nucleotide exchange factor that may function as an effector of RAC1 small G-protein.
The N-terminal domain of yeast ECM25 protein has been identified as containing a lipid binding CRAL-TRIO domain.[3]
Structure
The Sec14 protein was the first CRAL-TRIO domain for which the structure was determined.[4] The structure contains several alpha helices as well as a beta sheet composed of 6 strands. Strands 2,3,4 and 5 form a parallel beta sheet with strands 1 and 6 being anti-parallel. The structure also identified a hydrophobic binding pocket for lipid binding.
Human proteins containing this domain
C20orf121; MOSPD2; PTPN9; RLBP1; RLBP1L1; RLBP1L2; SEC14L1; SEC14L2; SEC14L3; SEC14L4; TTPA;
References
- ↑ Min KC, Kovall RA, Hendrickson WA (December 2003). "Crystal structure of human alpha-tocopherol transfer protein bound to its ligand: implications for ataxia with vitamin E deficiency". Proc. Natl. Acad. Sci. U.S.A. 100 (25): 14713–8. doi:10.1073/pnas.2136684100. PMC 299775
 . PMID 14657365.
. PMID 14657365. - ↑ Panagabko C, Morley S, Hernandez M, et al. (June 2003). "Ligand specificity in the CRAL-TRIO protein family". Biochemistry. 42 (21): 6467–74. doi:10.1021/bi034086v. PMID 12767229.
- ↑ Gallego O, Betts MJ, Gvozdenovic-Jeremic J, et al. (November 2010). "A systematic screen for protein-lipid interactions in Saccharomyces cerevisiae". Mol. Syst. Biol. 6 (1): 430. doi:10.1038/msb.2010.87. PMC 3010107
 . PMID 21119626.
. PMID 21119626. - ↑ Sha B, Phillips SE, Bankaitis VA, Luo M (January 1998). "Crystal structure of the Saccharomyces cerevisiae phosphatidylinositol-transfer protein". Nature. 391 (6666): 506–10. doi:10.1038/35179. PMID 9461221.
External links
- UMich Orientation of Proteins in Membranes families/superfamily-129 - Calculated spatial positions of CRAL-TRIO domains in membrane